BACTERIAL COLONY DETECTION
An AI-powered solution for precise segmentation and automated counting of bacterial colonies in microbiological lab analyses, developed in collaboration with FH Münster.
THE PROBLEM
Analyzing bacterial growth on petri dishes is fundamental to microbiology, but manual colony counting is time-consuming and tedious. In collaboration with FH Münster, I developed an automated solution that significantly accelerates the analysis process.
DATA PIPELINE
We built a specialized photo apparatus using a Raspberry Pi High Quality Camera that captures 9.24-megapixel images of petri dishes with bacterial cultures, providing a strong foundation for training machine learning models.
For efficient data annotation, I developed a custom web application integrating the Segment Anything Model (SAM). This allowed researchers at FH Münster and me to quickly and precisely create segmentation masks through simple click interactions, significantly reducing annotation effort. This produced a dataset of 200 precisely annotated images.
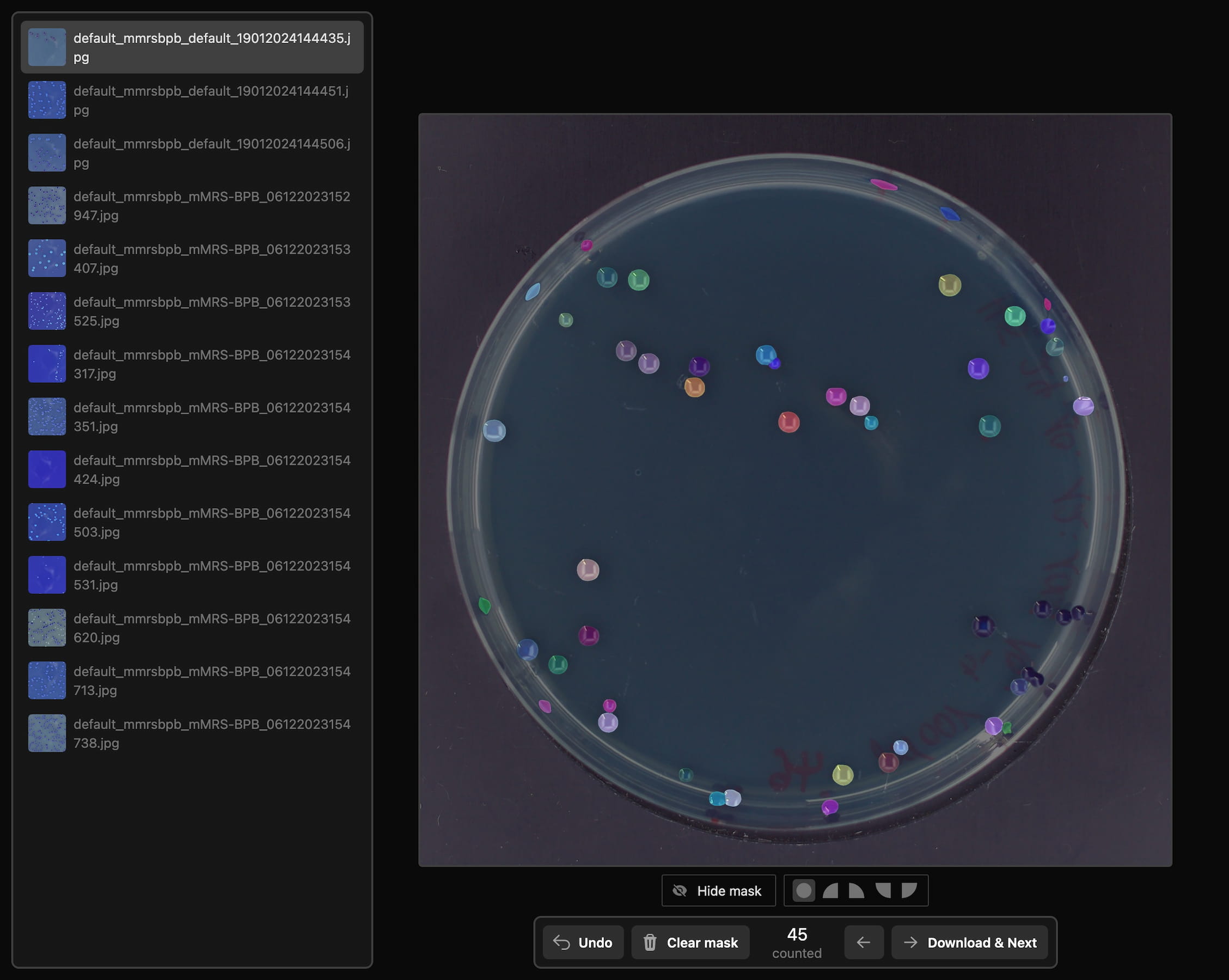
Precise instance segmentation masks created with the SAM-based annotation tool
MODEL TRAINING
I used the Faster R-CNN architecture, known for its high accuracy in object detection tasks. The model was strategically pre-trained on the COCO dataset, then fine-tuned on the specialized AGAR dataset, before final training on our custom-collected data. This multi-stage approach significantly improved model performance, achieving high precision with a low error rate.
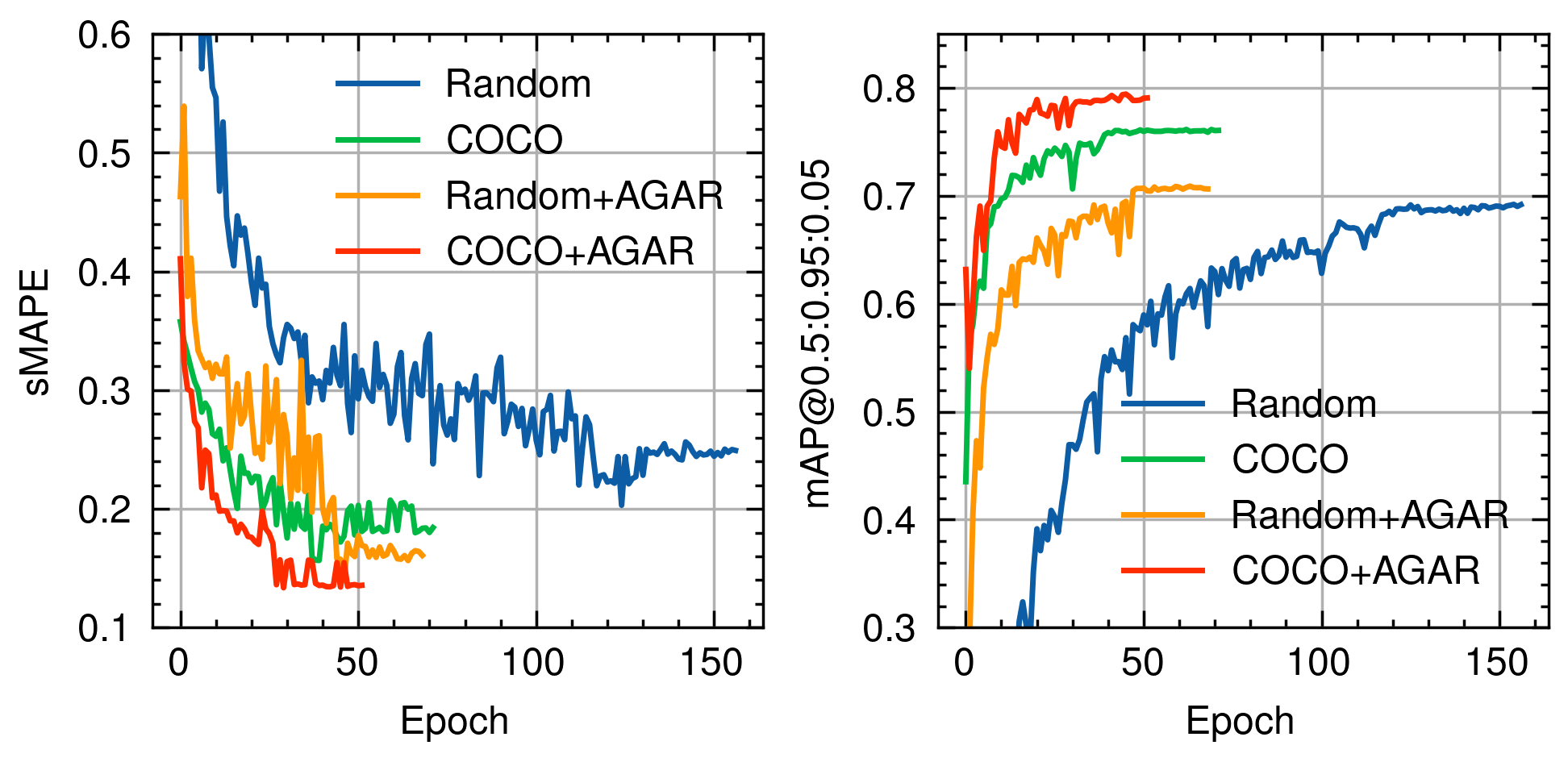
Validation performance improvement through strategic pre-training
RESULTS
The trained model detects bacterial colonies in approximately four seconds on a standard computer — a process that takes several minutes when done manually. The mean absolute error of 6.26 colonies per plate (out of often hundreds) demonstrates the model's robustness.
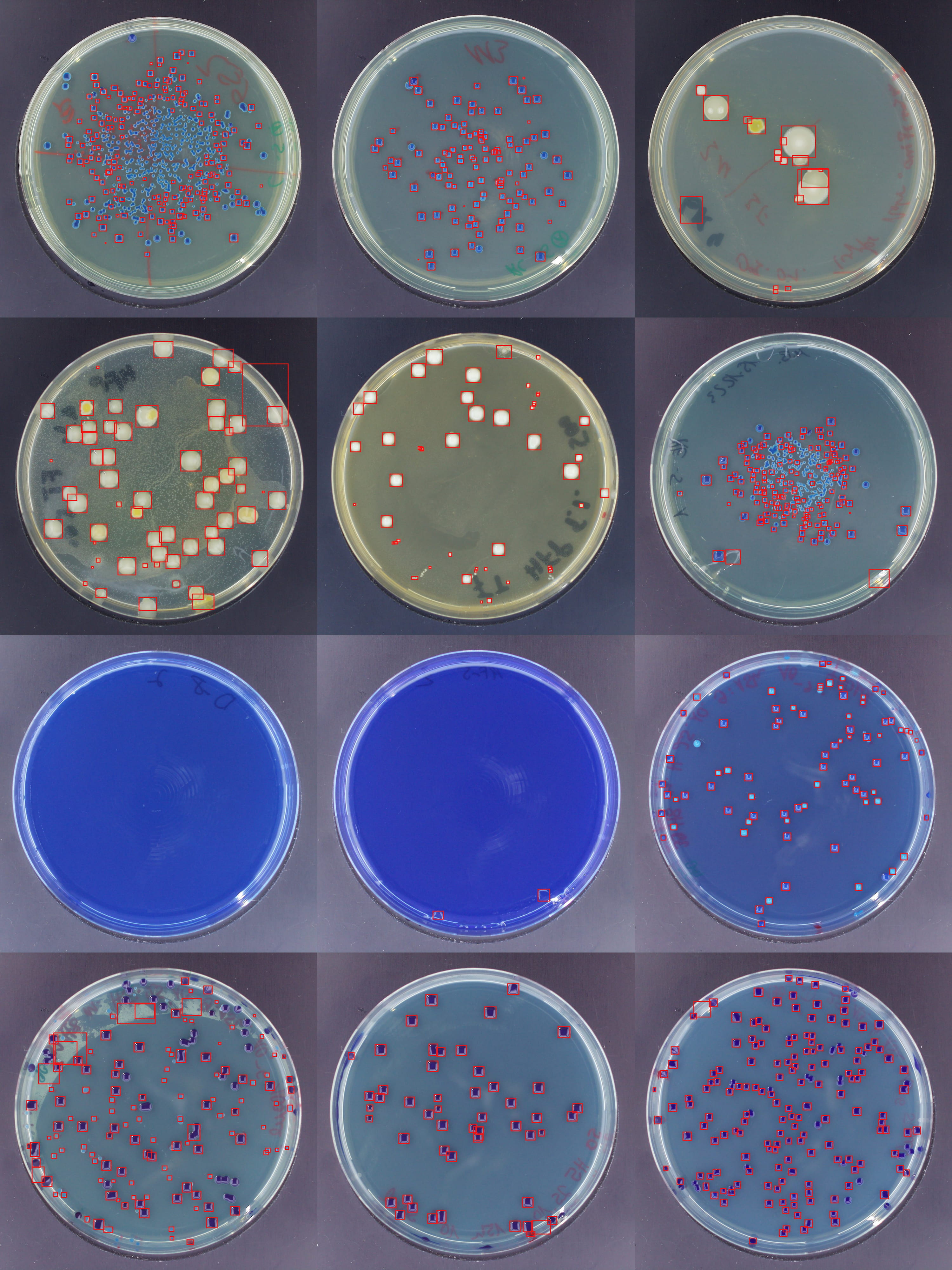
Detection examples across various colony densities and shapes
DEPLOYMENT
To ensure practical usability, I integrated the model into a user-friendly web application that is currently deployed at FH Münster, enabling researchers to quickly automate colony counting. The system is designed for scalability, with ongoing improvements possible through additional annotated data.
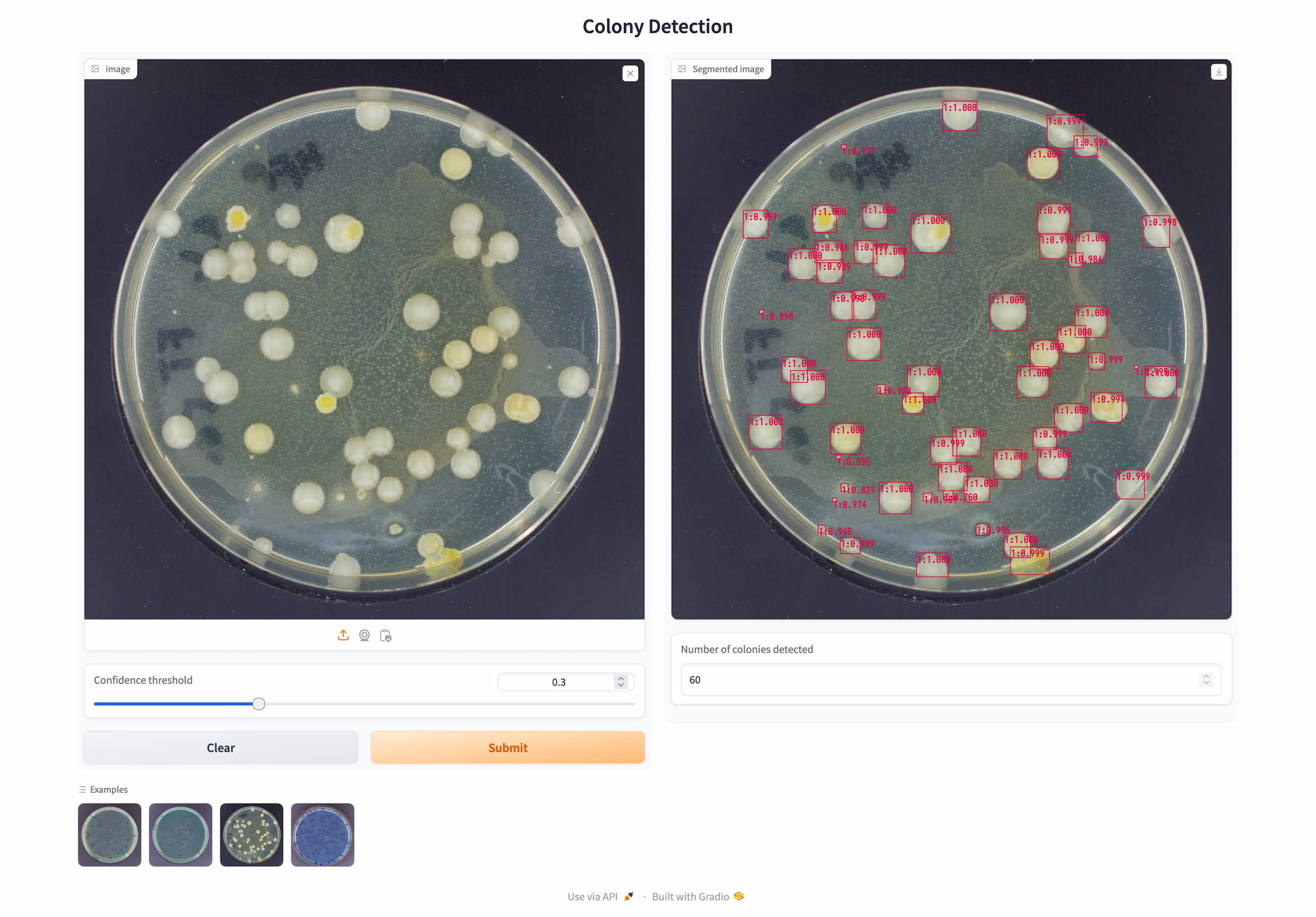
User-friendly inference application for quick colony counting